Supported A I Discovery Infrastructure for Engineering
Current Reservations
| Research Group | Reservation Time |
|---|---|
| Volkens Research Group | Till May 5, 2026 |
| Bradley Research Group | Till May 5, 2026 |
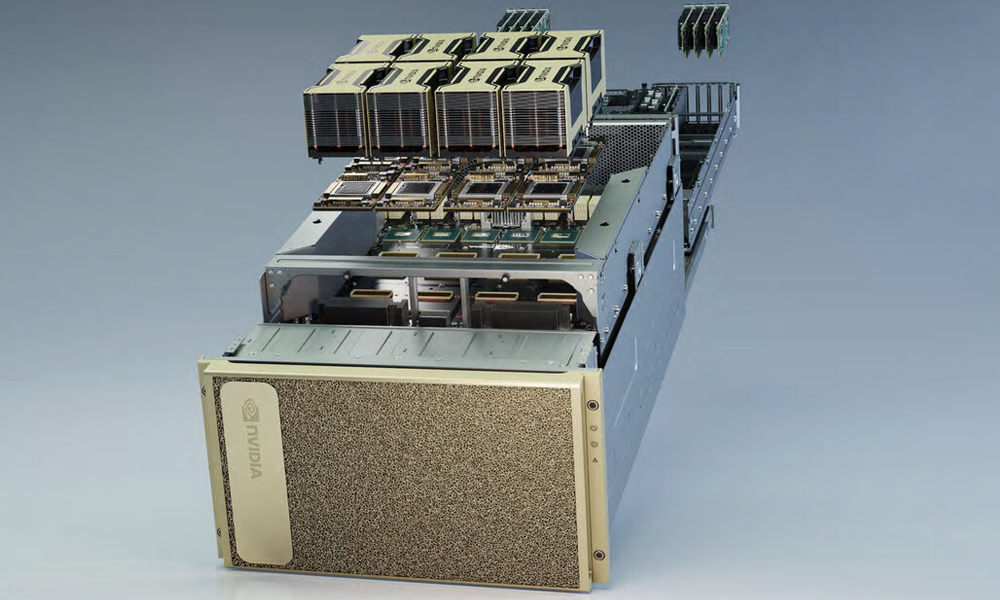

Cluster Machine List and Queue overview
Click here for a list of names and specs for each of the nodes in the cluster along with an overview of the queues that you can submit jobs to.
Getting Access and Eligibility.
The Saidie cluster is restricted to AI faculty and their groups. Additional WSCOE faculty performing research with AI may also be elligible. Fill out a freshservice request here and your request will be reviewed by The College Research Dean with the ETS Saidie administrators.
At this time while ETS is learning the admin and reporting tools and modifying the Saidie usage charge model, use of the machine is free. We expect a charge model to be in place by the end of summer 2026.
General Guidelines
- Use the job scheduler.
- Do not run jobs on the login node. Processes that impact the performance of the login node will be terminated without notice. If you need help running your job, please contact ETS.
- Be good to each other.
- This cluster is shared by several research groups. Please do your best to be fair and kind to others.
Partitions
Partitions are separate queues for submittied jobs and can contain overlapping groups of nodes. When you submit your job, resources are allocated from the partition’s nodes and your job runs on one or more of the nodes in that group. You can use the command sinfo or overview to see the list of partitions you can submit to. Below is a list of basic partitions configured on the Saidie cluster.
general- Use this partition for multiple short term jobs, Try and put in checkpointing for your program in case you need to restart it.
- This partition is capped at a max of 1 day of run time.
- This is the default partition
long-runs- This uses the same DGX/resources as the general partition.
- This partition is capped at a max of 7 days of run time.
Storage on the Cluster
Each user receives 2TB of space in the /home directory on Saidie’s all-flash storage server.
The storage on Saidie is a VAST solid state system and totals around 300TB. Users who require additional space may work with ETS to lease storage on a per-month basis according to the storage charge model at the bottom of this page.
Please note that this additional storage is short-term unlike the T:\Projects storage.
Research Groups may also add separate storage servers to the cluster which would be available under the root (/) directory. Contact us for more information.
Please contact us to purchase more individual space or find out about more storage options.
Running a Job On the Cluster
You can view the general getting started guide in pdf.
When working with a cluster you will not be running programs as you would on a personal computer or server. Instead, you interact with the cluster by issuing commands to the job scheduler. This is done via the command line with scripts submitted to the scheduler to run your job. A typical workflow would involve the steps below.
- Move your data/code to the cluster folder.
- For Linux and MacOS users, we recommend scp or sftp
- For Windows we recommend WinSCP
- Write or edit your submission script to add your required scheduler options and the commands needed to launch your program.
- Load any modules, activate virtual environments, and set any environment variables that are required; these are not added by default.
- Submit your job to the cluster with the “sbatch” command.
- e.g.
sbatch sample_submission.sh - Information on how to submit, watch, cancel, or find the status of your job please see the basic usage guide.
- e.g.
- Instead of outputting to your terminal window, by default standard output and errors from your job will be put in a text file in the same directory from which you submitted the job. You can then check any output or move it to another location for further processing. The location can be modified by SBATCH options.
Example Workflow
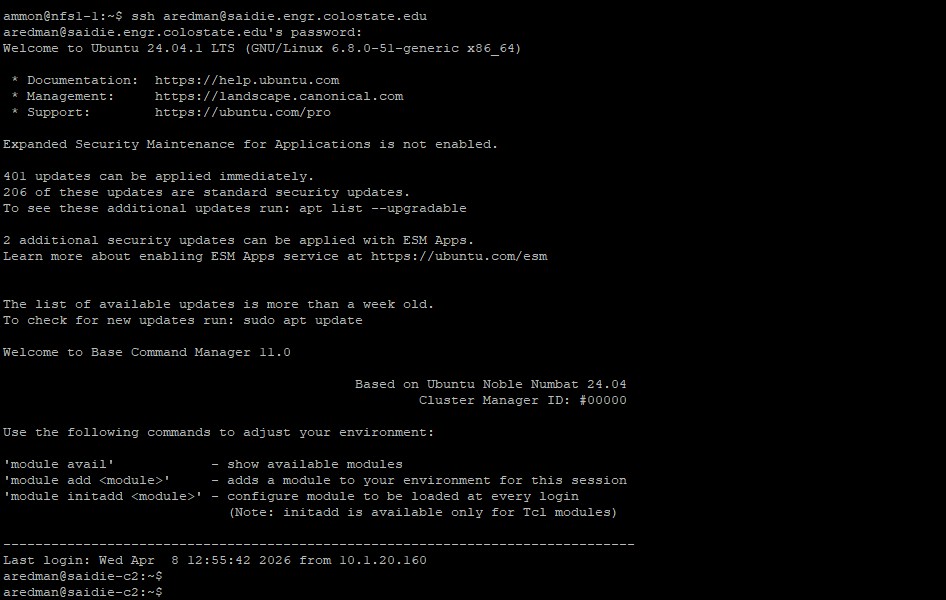
From your client machine, upload your data and connect to the cluster.
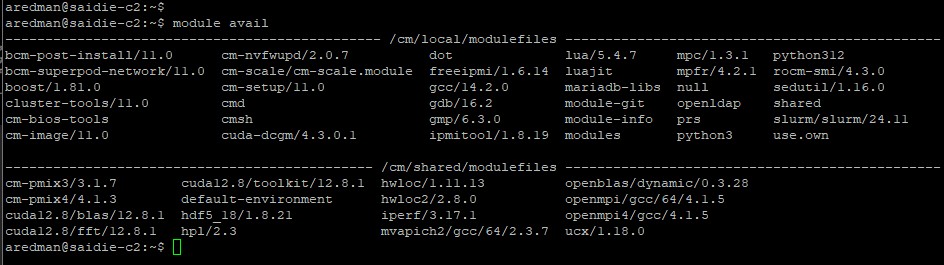
Load any needed modules.
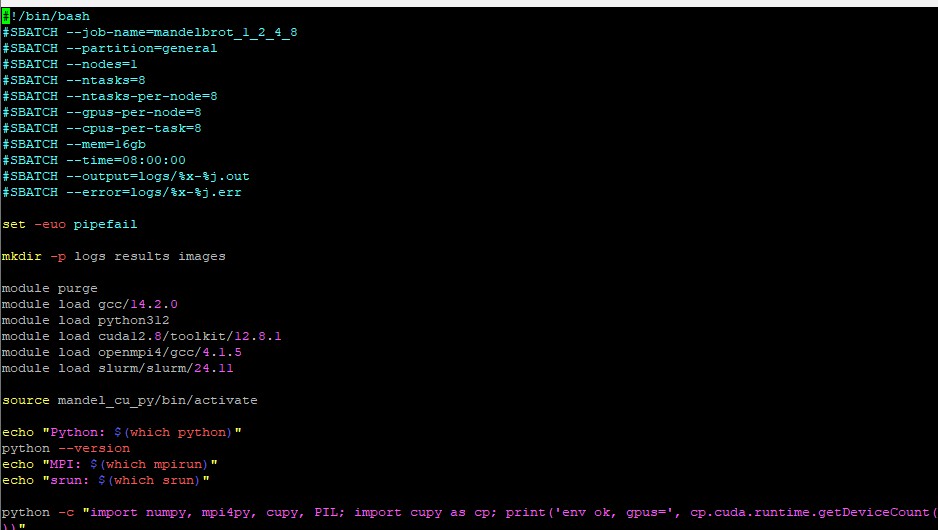
Edit your submission script. Here is a sample script to give you an idea of what you need. Your script will look different. The important flags are at the top with SBATCH. Make sure you include the --gpus, --cpus and --mem flags in your script.
Submit your job. Make sure to take note of your job number.

When your job is complete, check your data and output file for errors or standard output.

Sample Output